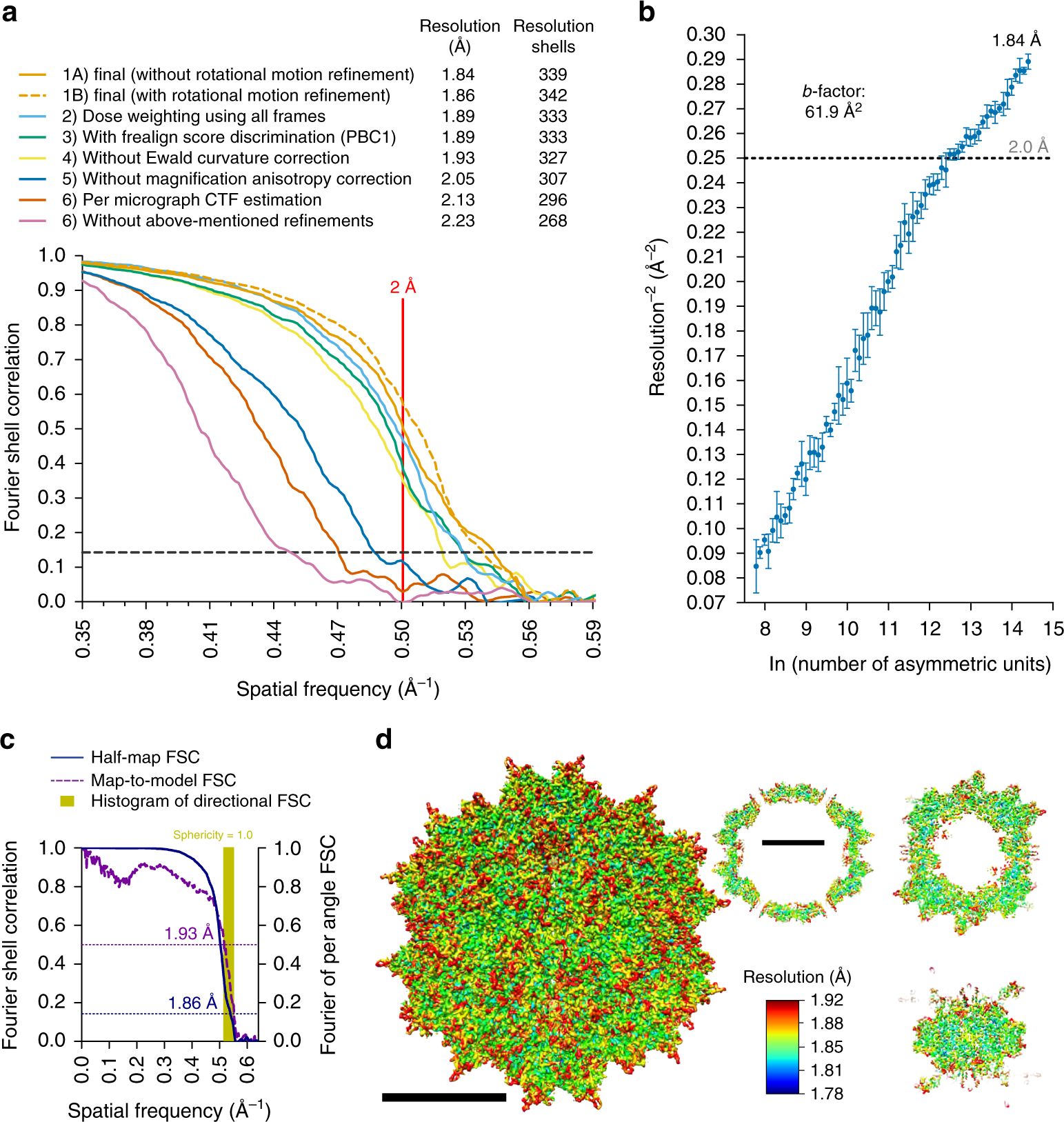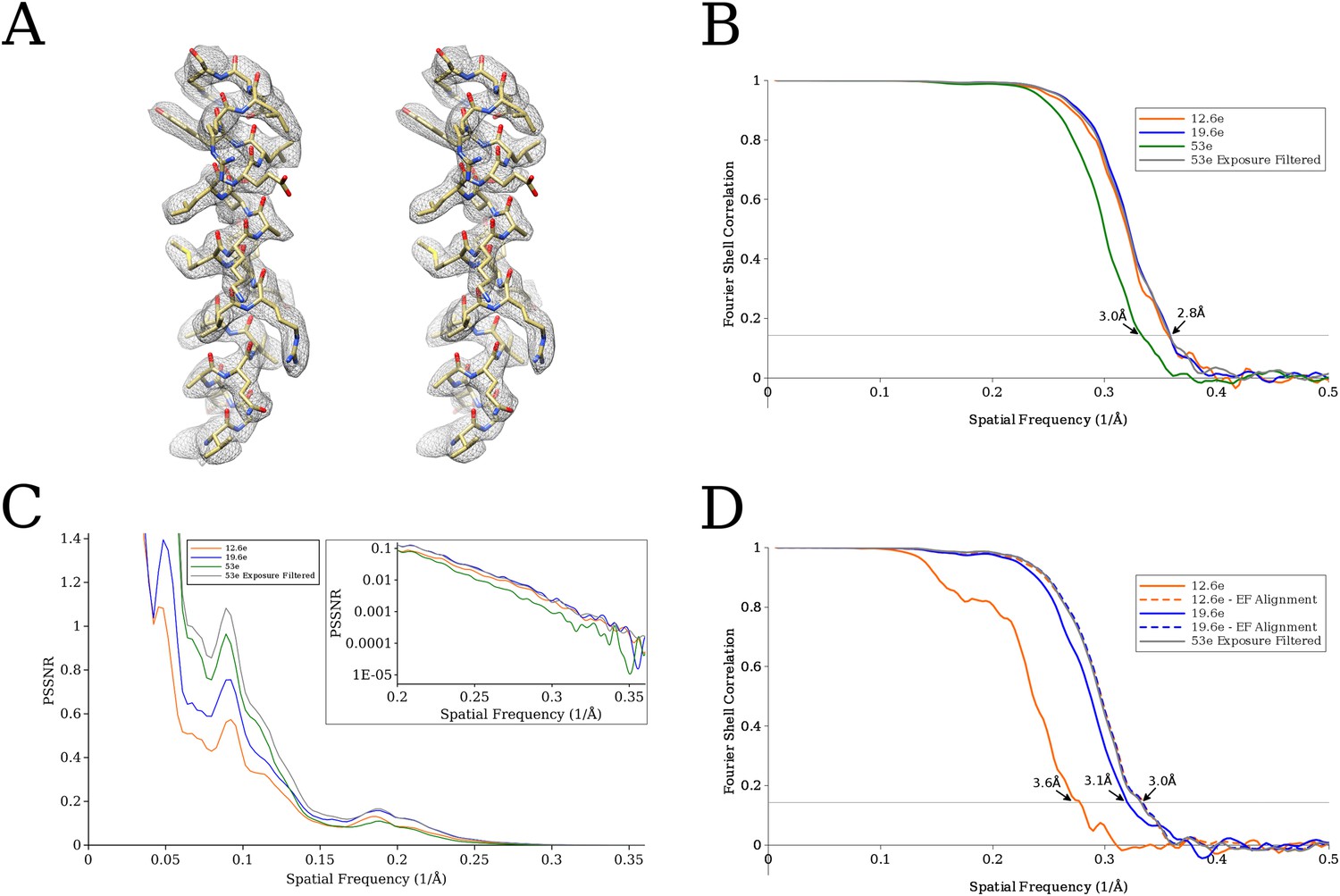
Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6 | eLife
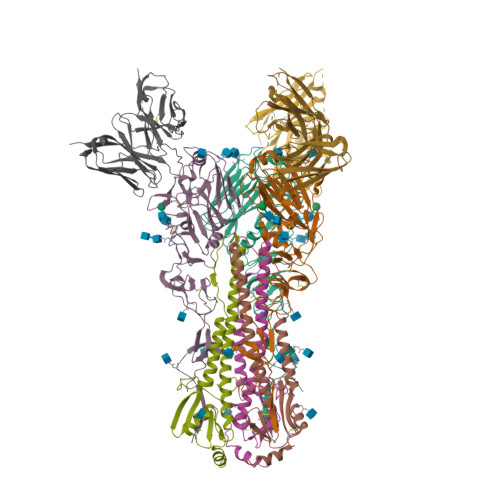
RCSB PDB - 5UK0: CryoEM structure of an influenza virus receptor-binding site antibody-antigen interface - Class 2

SBGrid: Structural Biology Software on Twitter: "Mark your calendars for next Tuesday. @SBGrid seminar by Johannes Elferich, @UMass, The Grigorieff Lab. High-throughput in-situ #CryoEM of hematopoietic differentiation https://t.co/QD5MIF2hIN https://t ...

RCSB PDB - 5UK0: CryoEM structure of an influenza virus receptor-binding site antibody-antigen interface - Class 2

Colleagues who have contributedt othe work described in this lecture.... | Download Scientific Diagram

From Electron Crystallography to Single Particle CryoEM (Nobel Lecture) - Henderson - 2018 - Angewandte Chemie International Edition - Wiley Online Library
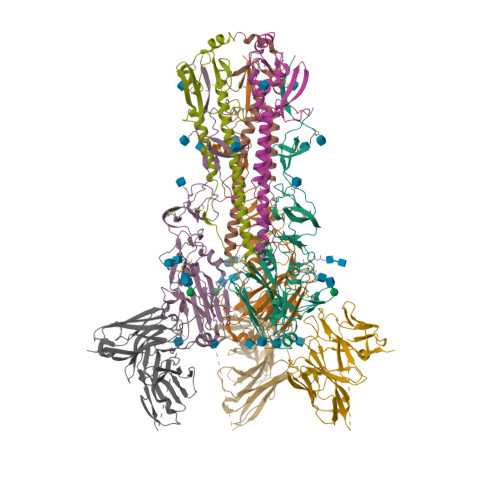
RCSB PDB - 5UK2: CryoEM structure of an influenza virus receptor-binding site antibody-antigen interface - Class 4